THE BRIEFING
Two companies independently beat Boltz-2 on binding affinity prediction - using completely different approaches. Isomorphic Labs unveiled a full drug design engine that goes well beyond AlphaFold. Terray Therapeutics ditched diffusion entirely and says it's 26x faster.
Meanwhile, OpenAI connected GPT-5 to Ginkgo's cloud lab and ran 36,000 experiments to cut protein production costs by 40%.
Arc Institute is scaling the perturbation data that virtual cell models need. And Phylo raised $13.5M to build a workspace where biologists orchestrate AI agents.
Let's dive in.
NEWS
OpenAI and Ginkgo close the loop

Still from a video released by OpenAI.
OpenAI connected GPT-5 to Ginkgo Bioworks' cloud laboratory - a robotic wet lab run remotely through software - and used it to optimize cell-free protein synthesis (CFPS), a method for making proteins without growing living cells. Over six rounds of closed-loop experimentation, the system designed and ran more than 36,000 unique reaction compositions across 580 automated plates, generating roughly 150,000 data points over six months.
The result: a 40% reduction in protein production cost ($422 per gram, down from the prior state-of-the-art $698) and a 27% increase in protein yield. GPT-5 found low-cost reagent combinations that human researchers hadn't tested - particularly formulations more robust in the low-oxygen conditions common in automated lab settings. It took the system three rounds and about two months to beat the benchmark.
Important caveats: this was demonstrated on a single protein (sfGFP). Generalization to other proteins and systems hasn't been shown yet. And it wasn't purely autonomous: human operators handled reagent preparation, loading, and fixing significant early variability issues. Every AI-designed experiment was validated against a programmatic model before execution to prevent hallucinated protocols.
Why it matters: This is the first published demonstration of a frontier model running a large-scale closed-loop biological experiment. The cost reduction is real and already commercial - Ginkgo is selling the improved reaction mix through its reagents store. But the single-protein limitation matters. The harder test is whether this approach generalizes across the many different biological processes that drug discovery actually requires.
Did you know? The AI-improved cell-free reaction mix can already be ordered over at Ginkgo.
NEWS
Isomorphic goes beyond AlphaFold with a full drug design engine

Isomorphic Labs - the Alphabet drug discovery company spun out of DeepMind - has unveiled IsoDDE, a unified computational drug design engine that goes well beyond structure prediction. The system more than doubles AlphaFold 3's accuracy on the hardest protein-ligand generalization cases, but the real news is what it does that AlphaFold never could.
IsoDDE is four tools in one. Structure prediction, yes - but also binding affinity prediction that surpasses all deep-learning methods and, remarkably, outperforms gold-standard physics-based approaches (FEP) on public benchmarks, without needing experimental crystal structures as a starting point. It predicts antibody-antigen structures at 2.3x the accuracy of AlphaFold 3 and 19.8x open source alternative Boltz-2 on challenging test sets. And it can identify novel druggable pockets on a protein from sequence alone, in seconds - approaching the accuracy of real-world fragment-soaking experiments.
A case study illustrates what this means in practice. Cereblon is a protein involved in tagging damaged proteins for disposal, as well as a key drug target. For 15 years, researchers thought cereblon had one principal drug-binding site. A 2026 study experimentally discovered a hidden cryptic pocket. IsoDDE recapitulated that discovery computationally, predicting both the known and the novel site using only the amino acid sequence as input.
Why it matters: This is the shift from “we can predict what a protein looks like” to “we can design a drug that binds to it and estimate whether it'll work.” Isomorphic is stacking capabilities - structure, affinity, pocket discovery, biologics - into a single engine. If the accuracy claims hold across real drug programs, this is what the AlphaFold investment was always meant to produce.
Did you know? Isomorphic Labs is hiring.
NEWS
Terray ditches diffusion - and beats Boltz-2
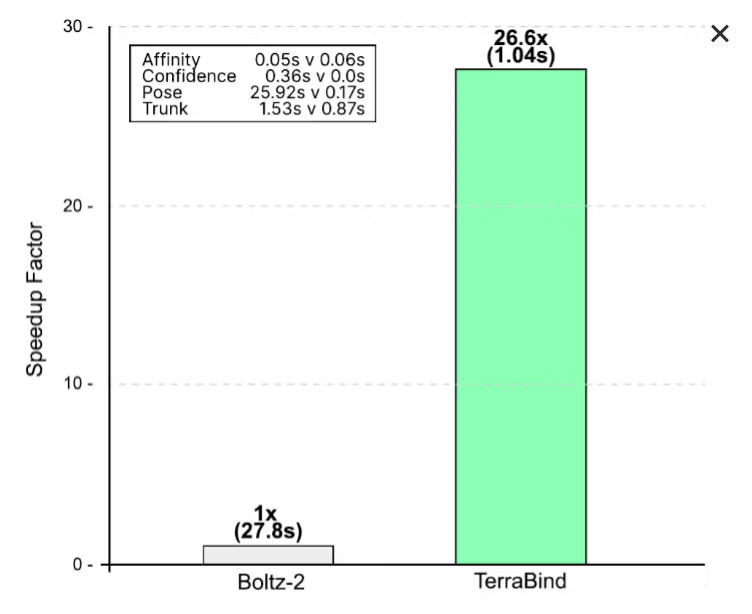
TerraBind is fast.
Binding affinity prediction just got a shake-up. Terray Therapeutics, a 140-person LA-area biotech, posted a preprint (as reported by Endpoints) showing its model EMMI Predict: TerraBind outperforming the widely used open-source Boltz-2 on binding affinity - 26x faster and roughly 20% more accurate.
The key move: Terray abandoned diffusion-based architectures - the approach behind AlphaFold and most leading structure models - in favor of a non-diffusion design that skips generating 3D structural visualizations entirely. The speedup is dramatic: 1.04 seconds per prediction versus 27.8 seconds for Boltz-2.
This puts Terray in an interesting race with Isomorphic Labs, which released its own Boltz-2-beating results days earlier (see our story above). On a shared CASP16 benchmark, Isomorphic scored 0.75 and Terray scored 0.73 (Boltz-2: 0.63–0.65). Both models are closed-source, making a full comparison difficult. But if Isomorphic is still using diffusion, Terray's speed advantage could be significant at screening scale - where you need to evaluate millions of candidates, not dozens.
The company, founded in 2018, raised a $120M Series B in 2024 and expects to select its first development candidate this year, with a clinic target of 2027.
Why it matters: The binding-affinity prediction race is heating up fast. Two companies independently beat the open-source benchmark in the same week, using different approaches. Terray's bet is that you don't need diffusion - or pretty 3D pictures - to screen drug candidates at scale. If the accuracy holds up, the 26x speed advantage matters when you're evaluating millions of molecules.
Did you know? Terray is hiring.
NEWS
Arc advances perturbation scaling for virtual cells

The sign in front of the Arc Institute building in Palo Alto, California.
Arc Institute has published new work on cellular perturbation data scaling laws.
Key finding: 4x cost reduction in perturb-seq (a technique for measuring how cells respond when you knock out specific genes) with 50% improved gene detection. They're partnering with Tahoe AI and Biohub to generate the largest open-source perturbation dataset for virtual cell models: 120 million cells.
Why it matters: Virtual cell models need massive perturbation data to simulate how cells respond to interventions. The data is the bottleneck. Making it cheaper and higher quality is infrastructure work that enables everything downstream - from drug target discovery to predicting side effects.
NEWS
Phylo Bio raises $13.5M for “agentic biology”

Phylo has announced its seed round.
Phylo launched with a $13.5M seed round co-led by a16z and Menlo Ventures' Anthology Fund (a fund created in partnership with Anthropic). The company grew out of Biomni, an open-source AI agent for biomedical research started at Stanford in 2024 - now used by over 7,000 labs, biopharma companies, and healthcare organizations.
The pitch: Biomni Lab as an “integrated biology environment” - a single workspace where biologists orchestrate AI agents across 300+ databases, software tools, and analytical systems. A case study with Ginkgo Bioworks showed it cut weeks-long cell-painting and transcriptomic workflows to hours.
Why it matters: This is a different bet than the OpenAI/Ginkgo approach. Ginkgo owns the lab infrastructure. Phylo is building software that sits on top of existing labs - potentially making agentic biology accessible to academic labs and smaller biotechs without billion-dollar capital expenditure.
Did you know? Biomni Lab is free to try at biomni.phylo.bio. They're also hiring and hosting launch events in South San Francisco and online.
THE EDGE
Derya Unutmaz - an immunologist who runs a research lab at Jackson Laboratory - built a database of 90,000+ driver mutations across 1,600 genes, filterable by cancer type. He built it using OpenAI Codex. No engineering team required.
The tool itself is useful if you work in oncology research. But the meta-point is worth noting: the barrier to building custom research tools has dropped dramatically. Expect more of these.
Until next time,
Peter at BAIO
