THE BRIEFING
AI drug discovery is scaling from individual models to full virtual organizations. Stanford spawned 37,000 AI agents to staff a virtual biotech that analyzed 56,000 clinical trials and surfaced a practical insight: drugs targeting cell-type-specific genes are 48% more likely to reach market.
Generate Biomedicines just raised $400 million in the largest biotech IPO since 2024 - betting public money that its AI-designed antibody can replace monthly asthma injections with twice-yearly dosing.
Bo Wang published both a case that diffusion models are better suited to biology than the architectures most foundation models currently use, and a review mapping where AI is reshaping every step of CRISPR.
And a YC startup open-sourced what it calls “Claude Code for biology” - a command-line agent wired to 190 drug discovery tools.
If there’s a through line here, it might be this: proof-of-concept is over. This issue is about infrastructure - organizational, financial and architectural, and open-source.
Let's dive in.
AD
Speak your prompts. Get better outputs.
The best AI outputs come from detailed prompts. But typing long, context-rich prompts is slow - so most people don't bother.
Wispr Flow turns your voice into clean, ready-to-paste text. Speak naturally into ChatGPT, Claude, Cursor, or any AI tool and get polished output without editing. Describe edge cases, explain context, walk through your thinking - all at the speed you talk.
Millions of people use Flow to give AI tools 10x more context in half the time. 89% of messages sent with zero edits.
Works system-wide on Mac, Windows, iPhone, and now Android (free and unlimited on Android during launch).
NEWS
Stanford builds a virtual drug company staffed by 37,000 AI agents
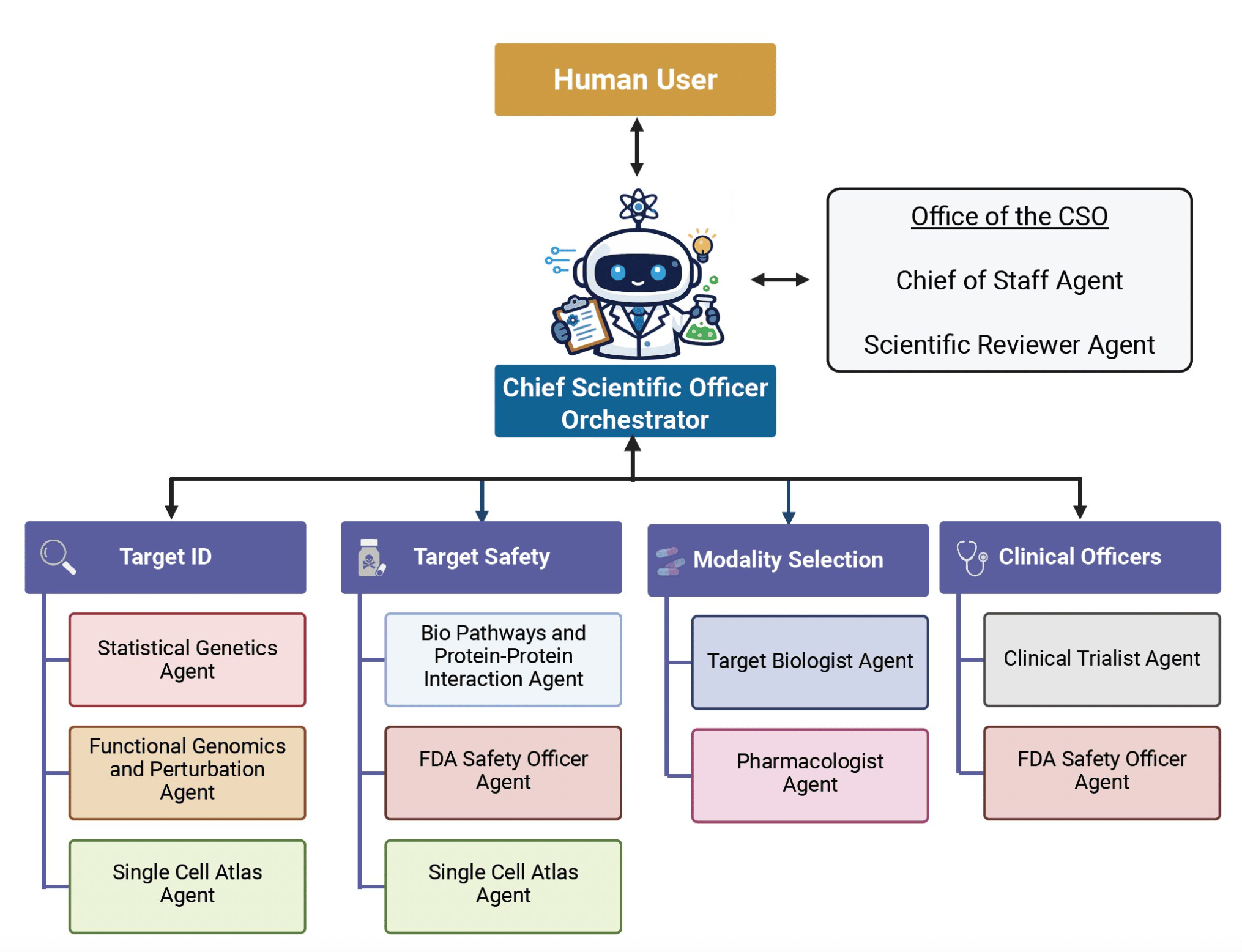
James Zou's lab at Stanford has built the Virtual Biotech - a multi-agent AI system that mirrors the structure of a pharmaceutical R&D organization. A virtual Chief Scientific Officer receives research questions, delegates to domain-specialized agents covering genetics, genomics, chemoinformatics, and clinical data, then synthesizes their outputs.
The headline demo: more than 37,000 AI agents annotated nearly 56,000 clinical trials and linked drug targets to multi-omic data including single-cell RNA sequencing. The finding - drugs targeting cell-type-specific genes were 48% more likely to reach market and showed 32% lower adverse event rates.
Two further case studies test the system on real questions: evaluating B7-H3 as a lung cancer target and analyzing a terminated ulcerative colitis trial to infer potential failure mechanisms and propose biomarker-guided enrollment strategies.
Why it matters: This extends Zou's Virtual Lab, published in Nature last year after its AI agents designed functional nanobodies for SARS-CoV-2. That system had a PI and a few specialists. The Virtual Biotech scales the concept to thousands of agents organized as a full R&D operation.
Did you know? The predecessor system - Virtual Lab - is open source on GitHub. You can create AI PI and scientist agents that hold research meetings to tackle scientific problems.
A WORD FROM OUR SPONSOR
The capital of longevity is back!

"We are living in an age where aging is becoming optional."
The key longevity event is happening in the Bay Area this May again.
Vitalist Bay (May 14–17) is a 4-day longevity festival where the longevity world actually converges — 100+ leading longevity speakers, 50+ hands-on workshops, 40+ activations to level up your health.
It's the place where the movement lives for four days and where you meet others likeminded taking longevity to the next level. Over 200 core people in the scene are already attending.
The first 15 BAIO readers to book get a 30% discount + complimentary DEXA scan with the code BAIO30 → https://vitalistbay.com/
NEWS
Bo Wang argues diffusion models could reshape biological foundation models
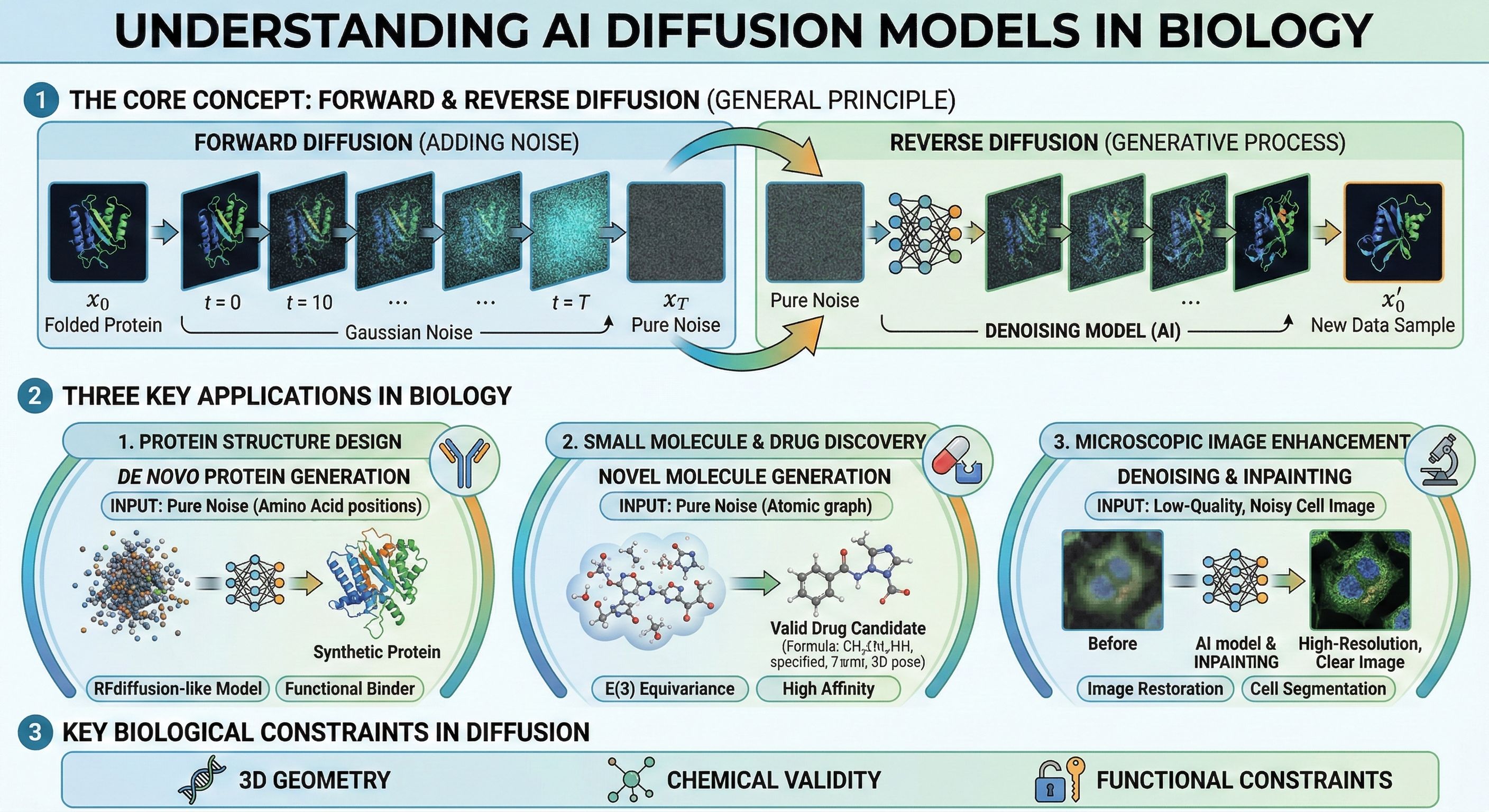
A Nanobanana 2 infographic showing how diffusion models work.
Bo Wang, the University of Toronto researcher behind EchoJEPA and scGPT, weighed in on a growing debate about AI architecture - and turned it toward biology. A blog post by ETH Zurich PhD student Dimitri von Rütte arguing that diffusion models will outscale today's dominant approach has been circulating widely.
Wang's take: the argument is actually stronger for biological foundation models than for language.
Some context on the two approaches. Most large language models today are autoregressive - they generate output one token at a time, left to right, each prediction conditioned on everything before it. Diffusion models work differently: they start from noise and iteratively refine the entire output at once, correcting mistakes at each step. Diffusion already dominates image generation (Midjourney, DALL-E) and protein structure design (RFdiffusion, Chroma).
Wang's point is that left-to-right generation loosely matches how sentences unfold, but has no basis in biology. A protein's function emerges from all its residues simultaneously. Gene expression is a high-dimensional state, not a narrative. DNA regulatory elements interact across hundreds of kilobases in both directions. The field already sensed this. ESM-2 - and its successor ESM-3 - use masked prediction rather than left-to-right generation, and ProteinMPNN already abandons strict left-to-right decoding. But many single-cell and genomic foundation models still inherit autoregressive architectures from NLP.
Why it matters: If the architectural mismatch is real, current bio foundation models may be leaving performance on the table. The debate reframes diffusion not as a niche technique for 3D structure, but as the natural paradigm for biological data broadly.
Did you know? Von Rütte's post cites evidence that diffusion models can train on the same data for many more epochs without overfitting - an advantage that could have particular relevance in biology, where high-quality training data is scarce, though neither author makes this connection explicitly.
NEWS
Generate Biomedicines raises $400M in largest biotech IPO since 2024
Generate Biomedicines, the company behind protein design model Chroma (also mentioned in the post above), raised $400 million in an IPO on the Nasdaq - the largest biotech listing since 2024. The company priced 25 million shares at $16, valuing it at roughly $2 billion. About $300 million will fund two Phase 3 trials of GB-0895, an AI-designed antibody for severe asthma, according to Fierce Biotech.
GB-0895 targets TSLP, the same pathway as AstraZeneca/Amgen's Tezspire - a drug that pulled in $1.2 billion in combined sales in 2024. Generate's ML platform engineered an antibody with roughly 20-fold higher binding affinity and a half-life of about 89 days versus Tezspire's 26, enabling six-month dosing instead of monthly injections. Phase 1 data presented at ERS 2025 showed a single dose suppressed inflammation biomarkers for at least six months. The Phase 3 program, SOLAIRIA, will enroll approximately 1,600 patients.
Why it matters: Public markets are now putting real money behind AI-designed biologics. The concrete differentiator here - six-month versus monthly dosing - came directly from computational optimization of binding affinity and half-life, a tangible output of generative biology. If Phase 3 confirms the design, patients would go from monthly injections to twice a year.
NEWS
AI is reshaping every step of CRISPR - here's the full breakdown

Bo Wang, who we mentioned above for his case for diffusion models in biology, has published a review in Nature Reviews Genetics mapping where AI is having measurable impact across the genome editing workflow.
The scope is broad. Deep learning models now predict on- and off-target activity for multiple CRISPR systems, from Cas9 to newer enzymes like TnpB. For base and prime editing, tools like PRIDICT predict editing outcomes from sequence context alone. Protein language models such as EVOLVEpro - which achieved up to 100-fold improvements in desired protein properties in a Science paper last year - are guiding directed evolution of Cas proteins to expand compatibility and reduce immunogenicity. And foundation models trained on metagenomic data are uncovering entirely new CRISPR systems from microbial diversity.
The section Wang highlights most: virtual cell models that could predict the consequences of any edit in any cell type - selecting targets, anticipating off-targets, modeling tissue-specific outcomes. The bottleneck, the authors note, is generating causally rich perturbation data at scale.
Why it matters: CRISPR and AI have mostly advanced on parallel tracks. This review argues they're converging - and that AI is becoming essential infrastructure for genome editing, not just a nice-to-have optimization layer.
Did you know? Wang's employer Xaira Therapeutics released X-Atlas/Orion last year - the largest publicly available Perturb-seq dataset, spanning 8 million cells across all ~20,000 human protein-coding genes. It's exactly the kind of perturbation data the review identifies as the bottleneck for virtual cell models. Available on Hugging Face. Oh, and Xaira is hiring.
THE EDGE
YC-backed CellType just open-sourced a command-line agent for drug discovery research - think Claude Code, but for biology. Install with pip install celltype-cli, type a research question in natural language, and the agent plans multi-step analyses using 190+ built-in tools spanning target prioritization, compound profiling, pathway enrichment, and safety assessment. It connects to 30+ database APIs - PubMed, ChEMBL, UniProt, Open Targets, ClinicalTrials.gov - with no setup required. Built on the Claude Agent SDK.
Until next time,
Peter at BAIO




