THE BRIEFING
Not every story in this issue is squarely AI × biology - but that's actually the point. To build virtual cells, predict perturbations, and design therapies computationally, the field needs infrastructure that doesn't always have “AI” in the title. A physics-based simulation of a complete cell cycle. A robot that produces a thousand nanoparticle formulations per hour. A discovery engine built by a DeepMind veteran and an MIT professor. These are the harnesses that make AI for biology possible.
The AI is here too: PerturbGen predicts how a single gene knockout ripples across a cell's entire future trajectory, validated in three biological systems. And Yann LeCun's $1 billion AMI Labs is betting that world models, not language models, are the right architecture for biological data.
Let’s dive in.
AD
AI Agents Are Reading Your Docs. Are You Ready?
Last month, 48% of visitors to documentation sites across Mintlify were AI agents—not humans.
Claude Code, Cursor, and other coding agents are becoming the actual customers reading your docs. And they read everything.
This changes what good documentation means. Humans skim and forgive gaps. Agents methodically check every endpoint, read every guide, and compare you against alternatives with zero fatigue.
Your docs aren't just helping users anymore—they're your product's first interview with the machines deciding whether to recommend you.
That means:
→ Clear schema markup so agents can parse your content
→ Real benchmarks, not marketing fluff
→ Open endpoints agents can actually test
→ Honest comparisons that emphasize strengths without hype
In the agentic world, documentation becomes 10x more important. Companies that make their products machine-understandable will win distribution through AI.
NEWS
A virtual cell, simulated from birth to division
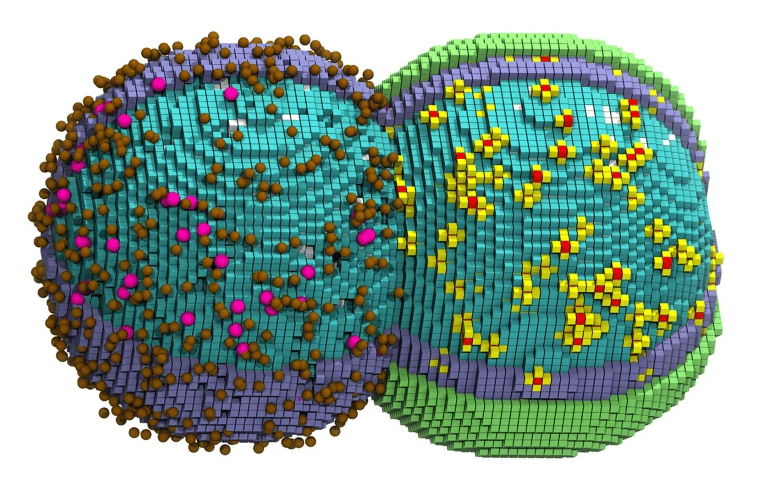
A computer generated illustration of a simulated cell in the early stages of division. Credit: Zane Thornburg/Cell
For the first time, researchers have simulated a living bacterial cell through its complete life cycle - from DNA replication to growth to division - while accounting for nearly every chemical reaction inside it. Zan Luthey-Schulten's team at the University of Illinois built a 4D model of JCVI-syn3A, a minimal bacterium with just 493 genes, and tracked its proteins, RNA molecules and metabolic reactions across the full 105-minute cell cycle at nanoscale resolution. The simulated cell divided within two minutes of the real thing. Published in Cell, each run took six days on a supercomputer.
Some details were simplified - water was approximated rather than individually modeled, and a few dozen genes with unknown functions were treated as inert placeholders. But the breadth is still unprecedented. This is physics-based simulation, not machine learning - yet it maps directly onto the virtual cell challenge. AI models aim to predict how cells respond to perturbations; this is the sort of mechanistic ground truth they need to validate against.
Why it matters: The gap between simulating 493 genes and the 20,000 in a human cell is vast, but a complete digital cell cycle - one that matches real-world timing - is now a concrete benchmark, and not just an aspiration.
Did you know? JCVI-syn3A was created at the J. Craig Venter Institute as one of the simplest self-replicating cells ever made. All simulation code is open source on GitHub.
NEWS
A $1 billion bet that world models, not language models, are the right architecture for biology

Yann LeCun left Meta last year and has now raised $1.03 billion for AMI Labs, a Paris-based startup building world models - AI that learns how physical systems behave rather than predicting the next word in a sentence. Biomedical research and drug discovery are among the target verticals. The direction is important: instead of generating outputs token by token, world models learn compressed representations of how things change over time.
Bo Wang, the University of Toronto researcher behind EchoJEPA (which we covered here) and scGPT, argues world models may fit medicine better than robotics. Medical images are full of noise unrelated to the patient. A language-model-style system tries to reproduce all that noise. A world model trained to predict what matters - like how a heart functions - can ignore it. Wang's EchoJEPA, trained on 18 million echocardiograms, needed just 1% of labeled data to reach 79% accuracy. Standard approaches hit 42% with all the labels.
Why it matters: A $1 billion bet that language models could be the wrong architecture for biological systems. Whether AMI delivers is years away, but the debate is already reshaping how biology AI gets built.
Did you know? AMI's CEO Alexandre LeBrun previously co-founded Nabla, a healthcare AI company that is now AMI's first disclosed partner. LeCun won the 2018 Turing Award for foundational work on deep learning. AMI is hiring across Paris, New York, Montreal and Singapore.
NEWS
A new AI model predicts how a single genetic change ripples through a cell's entire future
Most AI models that predict how cells respond to perturbations work on snapshots - what does the cell look like right after you knock out a gene? Mo Lotfollahi's lab at the Wellcome Sanger Institute and University of Cambridge asks a harder question: how does a perturbation reshape a cell's entire future trajectory? Their new model, PerturbGen, pre-trained on 107 million single-cell transcriptomes, predicts how genetic changes propagate across downstream cell states over time. The findings are published in a preprint and so has not been peer reviewed.
In the preprint, the authors report validation across three biological settings. In lab-grown skin tissue, it predicted which treatment would push cells toward a more natural state. In the blood system, it simulated a rare inherited disorder and matched the patterns seen in patient cells. In an immune response, it predicted how removing one inflammation gene early would reshape immune behavior hours later.
Why it matters: Genetic interventions are usually tested one cell state at a time. PerturbGen screens thousands of interventions across entire developmental or disease trajectories computationally - before anyone touches a pipette.
Did you know? All perturbation atlases are explorable at cellatlas.io/perturbgen. Lotfollahi pioneered single-cell perturbation prediction with scGen in 2019. Code is on GitHub.
NEWS
A robotic platform produces 1,000 lipid nanoparticle formulations per hour to feed AI models
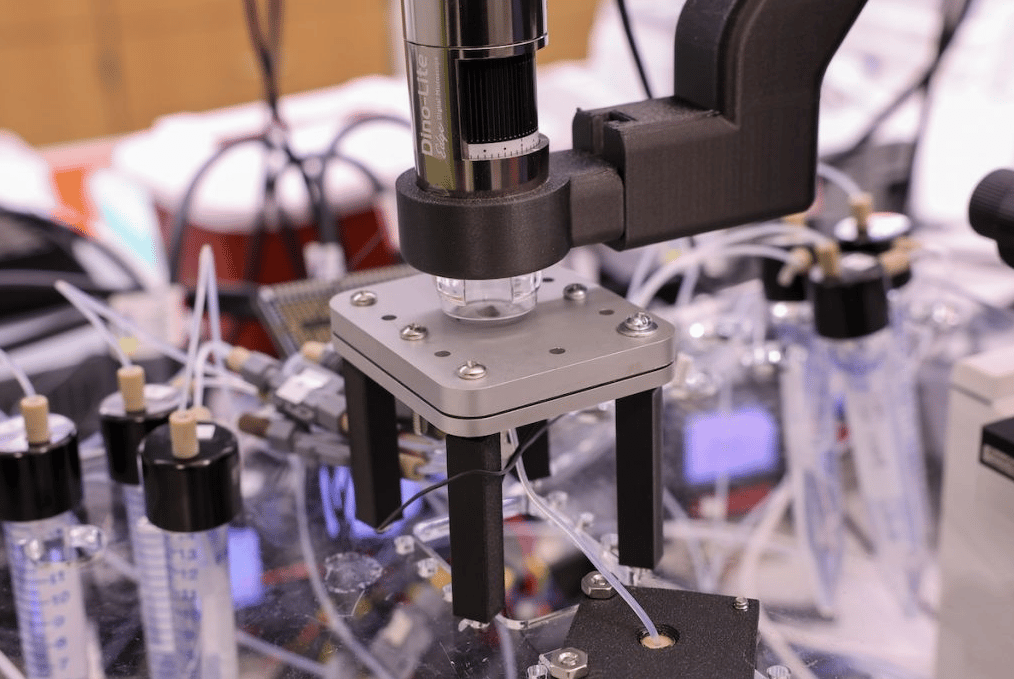

The design space for lipid nanoparticles - the delivery vehicles behind mRNA vaccines - spans roughly a quadrillion possible formulations. AI could navigate that space, but there isn't enough data to train on. The bottleneck: actually making the particles. Penn Engineering's Michael Mitchell and David Issadore built LIBRIS, a robotic microfluidic platform that produces around 1,000 LNP formulations per hour - roughly 100 times faster than conventional methods. The platform has been published in ACS Nano.
The machine uses a glass chip with parallel channels that mix lipid components under controlled pressure, creating up to eight distinct formulations simultaneously. Because the channels clean rapidly, it runs more or less continuously. The explicit goal is generating the large, systematic datasets that machine learning models need to predict how a particle's chemistry maps to its biological behavior.
Why it matters: mRNA delivery still relies too much on trial and error. LIBRIS is infrastructure for making that process computational - not by building the AI itself, but by producing the training data the field currently lacks.
Did you know? LIBRIS stands for LIpid nanoparticle Batch production via Robotically Integrated Screening. Mitchell's lab at Penn has been a leading group in LNP engineering for mRNA delivery.
NEWS
A DeepMind veteran and an MIT professor launch an AI discovery engine with $13.5 million

A Google DeepMind veteran and an MIT professor who has spent decades applying AI to biological materials have launched a startup to build what they call an AI discovery engine - a system designed to generate novel scientific hypotheses, not just retrieve existing ones. Unreasonable Labs raised $13.5 million in seed funding led by Playground Global, with Yuan Cao (senior research scientist on Google's Gemini team) as CEO and Markus Buehler (MIT, 500+ papers on AI-driven materials and protein design) as CTO. Robert Langer, the MIT biotech pioneer behind 40+ companies, advises.
The pitch: current AI models summarize what's known but can't compose genuinely new insights across disciplines. Unreasonable pairs large language models with what the founders call neurosymbolic abstractions - structured representations of physical laws and cross-domain relationships - to reason beyond training data. The company has initiated pilot collaborations in pharma, materials science and energy.
Why it matters: If the thesis holds, this is a general-purpose tool for any scientist doing discovery work - biologists included.
Did you know? Unreasonable is hiring in Palo Alto and Cambridge.
THE EDGE
The Stanford-Princeton team behind LabOS (which we covered here) has open-sourced LabClaw - 206 agentic skills that turn any OpenClaw agent into a biomedical research assistant. Skills span biology and life sciences, drug discovery, clinical research, data science and literature search, composable into end-to-end research workflows. One message to your OpenClaw agent installs the full library. Code on GitHub.
ON OUR RADAR
Until next time,
Peter at BAIO