THE BRIEFING
Three stories in this issue share a pattern: the AI caught something that standard methods missed.
A clinical trial superforecasting system predicted a drug would fail - before the results came in - by spotting a design flaw the company had overlooked. A blood proteomics model predicted who would decline cognitively faster than the diagnoses doctors had given. And a single-cell model that refused to compress biology into one number surfaced a drug target that conventional analysis would have buried.
Meanwhile, three veterans of Ginkgo Bioworks launched a design studio arguing that biology's real bottleneck isn't better models - it's a missing design language. And the Stanford-Princeton team behind LabOS built a simulation world where AI lab agents can practice experiments.
Let's dive in.
AD
How 2M+ Professionals Stay Ahead on AI
AI is moving fast and most people are falling behind.
The Rundown AI is a free newsletter that keeps you ahead of the curve.
It's a free AI newsletter that keeps you up-to-date on the latest AI news, and teaches you how to apply it in just 5 minutes a day.
Plus, complete the quiz after signing up and they’ll recommend the best AI tools, guides, and courses — tailored to your needs.
NEWS
An AI predicted a clinical trial would fail - before the results came out

Rohil Badkundri - a co-author on the ESM3 paper at EvolutionaryScale - has launched Warpspeed, an AI superforecasting system designed to predict clinical trial outcomes. Built through his 76th Street Research, the system uses what Badkundri calls a fleet of autonomous research agents, spending thousands of dollars in compute per forecast, to build a statistical model that simulates a trial's likely outcome.
To make the case, Badkundri spotlights Gossamer Bio. The company ran a Phase 3 trial of an inhaled drug for a serious lung condition. When the results were announced in late February, the drug had improved patients' walking distance by 13.3 meters, not enough to clear the statistical bar the company had set. The stock dropped about 80%. Warpspeed had estimated a 29% chance of success and predicted roughly 10 meters - close to what actually happened. Badkundri's argument: the failure was predictable from the trial's design. The company powered the trial expecting three times the effect that similar drugs have historically delivered.
In total, Badkundri says the Warpspeed system predicted five trial outcomes before their results came in - getting four right. All five were published together on March 30, after the results were known. The team is upfront about this: “We didn't cherry-pick, but feel free to take them with a grain of salt.” The real test starts now. Ten new predictions are live, with several trials reporting in weeks.
Abhishaike Mahajan, aka owl_posting, noted that Warpspeed's one miss is telling - the system defaulted to what similar drugs typically do and missed that this particular molecule would outperform its class. For now, Mahajan argues, the system works best as “what a very careful analyst would conclude with infinite patience and perfect recall of the literature.” Whether it can go beyond that - predicting outcomes for genuinely novel drugs where there's no class history to anchor to - is the open question.
Why it matters: The bigger thesis is about what's broken in biotech. Trials fail all the time for reasons that have nothing to do with whether the drug works - wrong patient selection, noisy endpoints, designs that assume an effect size the drug can't deliver. That makes the entire ecosystem risk-averse. Badkundri points to history: Pfizer walked away from GLP-1 in the late 1980s despite compelling data. Investors compared mRNA startups to Theranos. Jim Allison spent years pitching cancer immunotherapy to companies that dismissed it. These drug classes went on to save millions of lives. If AI superforecasting can flag predictable failures before hundreds of millions are spent, the argument goes, sponsors gain the confidence to take bigger swings on biology instead of crowding into the same safe hypotheses.
Did you know? Warpspeed isn't the only player. Insilico Medicine's inClinico system, for example, has reported 79% accuracy on prospective trial predictions. All ten of Warpspeed's new forecasts - with adjustable statistical models - are live at warpspeed.sh.
NEWS
An AI that refuses to compress biology into a single number found what standard analysis missed
Most AI models for single-cell biology compress everything about a cell into one summary - a single mathematical representation that captures its overall state. Mo Lotfollahi's group at the Wellcome Sanger Institute and University of Cambridge argues that this loses important detail. Their new model, Tripso (Transformers for learning Representations of Interpretable gene Programs in Single-cell transcriptOmics), instead gives each biological program its own separate representation - one for a signalling pathway, another for a transcription factor's targets, and so on. The result is a decomposed view of what a cell is doing, program by program. The paper positions it as a foundation for “interpretable virtual cell models” - a direct contribution to the race we've been tracking with PerturbGen (Issue 7, also from Mo Lotfollahi's lab) and X-Cell (Issue 9).
The strongest demonstration: Tripso compared lab-grown blood stem cells to their counterparts in the human body, viewing them through multiple program lenses. Two different programs - one for a growth signalling pathway, one for low-oxygen response - independently flagged two different genes that were more active in cells drifting away from a stem-like state. Both genes turned out to be parts of the same protein assembly line in the cell - the SEC61 translocon - a convergence that conventional gene-by-gene analysis would have missed, because the individual effects were too small to stand out among thousands of other genes. When the team inhibited SEC61 in culture, stem cell frequency rose from 1.4% to 5.0%.
Beyond stem cells, Tripso revealed age-specific patterns in blood development and discovered a previously unknown immune cell program linked to eczema - confirmed by mapping gene activity and protein expression directly in tissue sections.
Why it matters: Lotfollahi pioneered single-cell perturbation prediction with scGen in 2019. The paper argues that collapsing a cell into a single representation hides the biology that matters most - the specific programs you'd want to target. The SEC61 result shows this isn't just theoretical: the program-level view surfaced a target that a gene-level view would have deprioritized.
Did you know? Tripso is open-source on GitHub. Lotfollahi is also scientific co-founder of AI VIVO and serves on the scientific advisory board of Novo Nordisk.
NEWS
AI diagnoses multiple brain diseases from a single blood draw
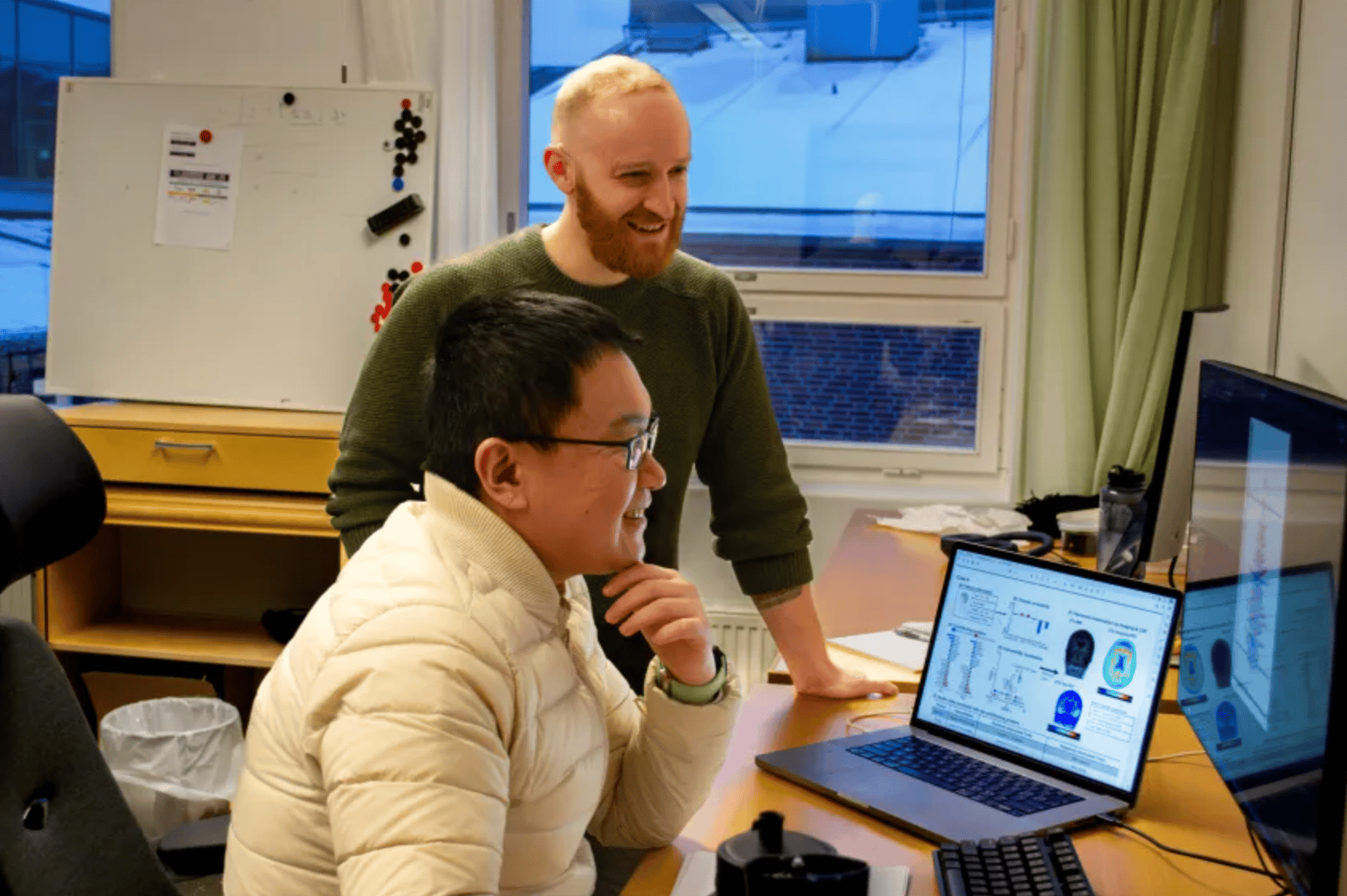

Two of the researchers behind the AI model, Jacob Vogel and Lijun An, show the results of their study. Credit: Emma Nyberg/Lund University.
A deep learning model trained on blood samples from more than 17,000 patients can simultaneously screen for Alzheimer's, Parkinson's, ALS, frontotemporal dementia, and prior stroke - from a single draw. Jacob Vogel and Lijun An at Lund University built ProtAIDe-Dx (Proteomics-based Artificial Intelligence for Dementia Diagnosis) on plasma proteomics data from the Global Neurodegenerative Proteomics Consortium - the largest dataset of its kind for neurodegenerative diseases. The model achieved 70-95% balanced accuracy across all five conditions plus healthy controls, with ALS (95%) and Parkinson's (92%) performing strongest.
The finding that may matter most: the model's predictions distinguished who would decline faster - and clinical diagnoses didn't. Across the dataset, the diagnoses doctors gave couldn't significantly separate rates of cognitive deterioration after statistical correction. The AI's protein-based predictions could, regardless of original diagnosis. Many patients diagnosed with Alzheimer's showed protein patterns more consistent with other conditions, suggesting misdiagnosis or co-existing diseases. Misdiagnosis rates in specialist dementia clinics run 25-30% and can exceed 50% in primary care.
The authors are candid about limits: plasma proteomics alone cannot yet replace current clinical markers like brain imaging and spinal fluid tests, and accuracy dropped substantially when the model was applied to hospitals and cohorts not included in training. Published in Nature Medicine.
Why it matters: Diagnosing which neurodegenerative disease a patient has is hard. Detecting that they might have more than one - which occurs in 70% of patients over 80 - is even harder. A blood test that simultaneously screens for multiple conditions and flags co-pathology could change how early these diseases are caught and how clinical trials select patients.
Did you know? Did you know? The GNPC dataset is accessible to qualified researchers at neuroproteome.org, and code is open-source on GitHub.
NEWS
Three ex-Ginkgo leaders launch a design studio for biology in the AI era

Christina Agapakis, Jake Wintermute, and Patrick Boyle - all formerly of Ginkgo Bioworks - launched American Wetware, a design studio for bioproducts. Their argument: biology has powerful new tools but still lacks a design language. The Bauhaus gave industrial manufacturing one. The GUI gave personal computers one. Biology never got its equivalent - and AI won't fix that on its own.
One of six planned offerings is a “test kitchen for AI scientists” - a sandbox wet lab where AI agents get tested against real biology. The others span bioproduct design, biotech marketing, content, research models, and an AI × bio policy think tank.
Why it matters: American Wetware makes a pointed case that neither academia (“it simply isn’t the role of academia to ship products”), nor pharma (“biopharma’s culture of secrecy means that no general community of practice can form to share ideas across projects”) will produce the design language biology needs.
Did you know? American Wetware’s manifesto previews some of the bioproducts the company suggest we might test: peptides that taste like chicken, wearable biosensors for THC, leather made from your own skin cells. “Do people want these things?” they ask. Finding out is the point.
NEWS
The LabOS team built a simulation universe for AI lab agents

Le Cong's lab at Stanford and Mengdi Wang's at Princeton - the team behind LabOS (which we covered in Issue 5), LabClaw (Issue 7), and MedOS (Issue 8) - released LabWorld, a simulation environment where AI agents practice running biomedical experiments. The system includes 100 realistic lab assets, 1,000 skills (the primitive actions like grasping, transferring, and dispensing that make up lab work), and over 10,000 executable biomedical protocols. NVIDIA is a founding partner. No preprint yet - just a project page and demos.
Why it matters: BAIO has tracked this team from smart glasses that watch scientists work, to an agent skill library, to hospital pilots. LabWorld adds another layer: a place where AI lab agents can fail cheaply and learn fast before running real experiments.
Did you know? You can explore LabWorld here.
THE EDGE
Adaptyv Bio officially released its wet-lab API - giving AI agents and protein designers programmatic access to Adaptyv's automated lab in Lausanne. Design a protein computationally, submit the sequences through a few lines of code, and the lab synthesizes it, runs binding assays, and returns results in about three weeks. Tamarind Bio and Phylo Bio (whose Biomni Lab we covered in Issue 1) have already integrated it. Co-founder Julian Englert says the company has tested tens of thousands of proteins over three years for pharma, AI labs, startups, and academic researchers.
ON OUR RADAR
Until next time,
Peter at BAIO



